buildAGL-Score Input
local_libraryReferences
1 Duc D Nguyen and Guo-Wei Wei, AGL-Score: Algebraic Graph Learning Score for Protein-Ligand Binding Scoring, Ranking, Docking, and Screening, Journal of Chemical Information and Modeling, 59, 3291-3304, (2019) Read
2 Duc D. Nguyen and Guo-Wei Wei, Algebraic graph learning of protein-ligand binding affinity (2018)
Read
local_libraryTraining Datasets
Below are the datasets used to train AGL-Score for docking and screening power assessments
- Docking Power Dataset: Poses folder contains the structures for training data, INDEX folder consists of labels of the corresponding structures in Poses folder
- Screening Power Dataset: Poses folder contains the computer-generated structures for training data and test data, INDEX folder consists of predicted energies of the training data in the Poses folder
perm_contact_calendarContact Information
Contact us : weig at msu.edu
Webmaster : ddnguyen at msu.edu
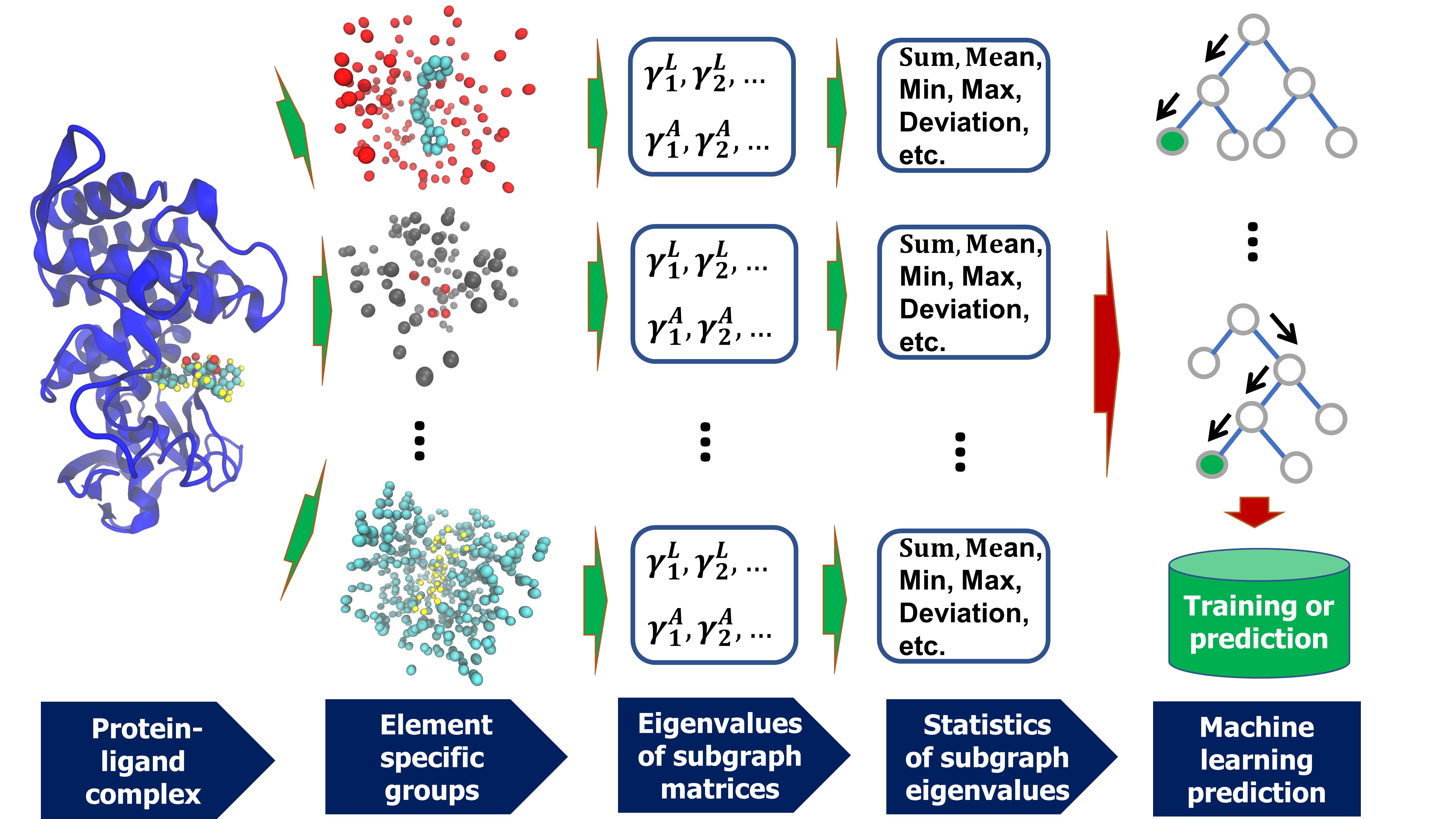
layersAGL-Score's Flow Chart